We first define:
- Genotype: Internal encoded form manipulated by GA operators
- Phenotype: Actual candidate solution in the problem domain
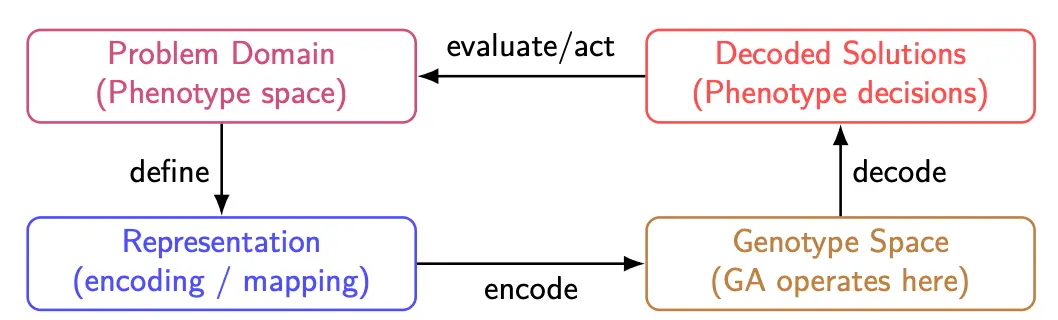
An example is genotype = 011101001, phenotype = 17.8.
Good genotypes induce smooth, meaningful neighborhoods in phenotype space.
Binary Representation
A standard way of doing encoding is with binary representation, where decision variables are represented as bit strings:
- Advantages: Simple and universal, well-defined crossover and mutation, theoretical tractability
- Disadvantages: precision requires long chromosomes, artificial discontinuities may appear
Binary encoding maps a bit string to a real decision variable. For a variable encoded using bits:
Resolution (precision):
Each bit string corresponds to a discrete point in . Increasing makes the grid finer. Thus, the genotype search space grows at . Higher precision improves resolution, but increases chromosome length and search complexity.
Hamming distance
The Hamming distance between two bit strings is the number of bit positions at which they differ. In binary encoding, a small Hamming distance does not guarantee a small phenotype change, as adjacent integers may differ by many bits.
- Phenotype distance:
- Hamming distance: 4
Gray coding
Gray coding is a representation designed to preserve neighborhood structure. Consecutive integers differ by exactly one bit, so Hamming distance better correlates with numeric distance.
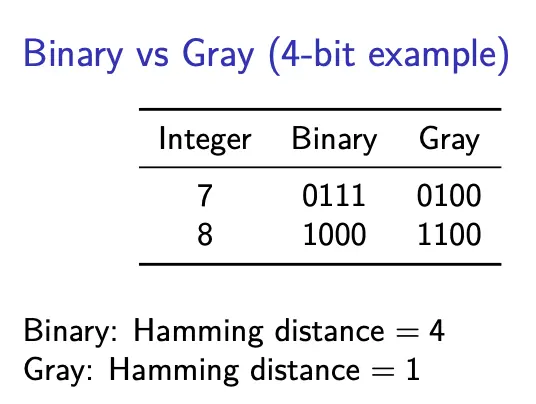

This results in a smoother genotype-phenotype mapping, in turn resulting in improved local search behavior.